Sr2 is a stem rust resistance gene that has been used in breeding for around 60 years as a source of durable and broad-spectrum adult plant resistance, which includes resistance to Ug99 and its related isolates. Sr2 is located on the short arm of chromosome 3B and confers partial resistance only in the homozygous state (recessive resistance gene). It was originally transferred from Yaroslav emmer wheat into hexaploid wheat.
Traditional breeding with Sr2 has been difficult because of its recessive nature and the fact that its phenotype is only evident at the adult plant stage and can be influenced by the genetic background and the environment. One partially dominant morphological marker linked to Sr2, pseudo-black chaff (PBC), has been used for years in breeding programs. PBC is a dark pigmentation that occurs on the glumes, peduncle and below stem internodes, but its levels of expression vary with genetic backgrounds and environments. In 2003 Spielmeyer et al (1) determined that a 120-bp allele of the microsatellite locus Xgwm533 was tightly linked to Sr2 in several lines. This marker is complicated to use because there are two different Xgwm533 on 3BS: several lines that did not carry Sr2 also amplified a 120-bp product for Xgwm533, which actually was another SSR allele that differed in sequence. Hayden et al (2) developed sequence-tagged microsatellites (STM) markers that could differentiate between the different Xgwm533 variants. Later, McNeil et al (3) using physically ordered BAC clones near Sr2 found three new SSR loci that mapped closer to Sr2 than Xgwm533 did, but they were quite polymorphic among lines with or without the resistance gene. Some of them could be useful for breeding but in any case a careful check of the parents is needed.
Also, a set of KASP markers was developed in 2013. The information on the primers needed is detailed at the bottom of this section below, along with the expected polymorphisms.
CAPS Markers for Sr2
Contributed by W. Spielmeyer, R. Mago, E.S. Lagudah, J. Ellis (CSIRO Plant Industry, Canberra, Australia)
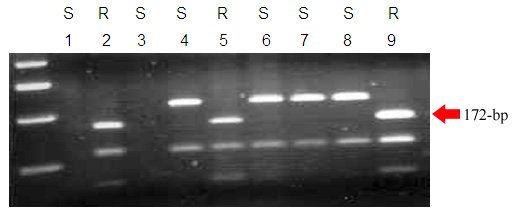
The CAPS marker presented here, csSr2, was derived from the Sr2 locus and detects three different alleles of Sr2 with a high accuracy (Fig. 1):
- ‘null’ allele or lack of amplification, associated with most Australian non-Sr2 wheats
- Marquis type allele: Two fragments (225 bp and 112 bp) after amplification and cutting with the appropriate restriction enzyme (BspHI). This allele is associated with non-Sr2 wheats prevalent in North America and CIMMYT.
- Hope type allele: Three fragments (172 bp, 112 bp and 53 bp) after amplification and cutting with BspHI enzyme. This allele is associated with the presence of Sr2.
Primers sequences:
csSr2-F 5'- CAA GGG TTG CTA GGA TTG GAA AAC -3’
csSr2-R 5'- AGA TAA CTC TTA TGA TCT TAC ATT TTT CTG -3’
PCR conditions:
- Denaturing step: 95°C, 2 min
- Amplification step (30 cycles):
- 95°C, 30 sec
- 60°C, 40 sec
- 72°C, 50 sec
- Extension step: 72°C, 5 min
- 15°C, 1 min
PCR mix and CAPS conditions:
- 2-4µl (100ng) DNA template
- 0.2mM dNTP
- 10 pmol of each primer
- 1.5mM MgCl2
- 1x GoTaq Flexi buffer
- 0.5 U GoTaq Flexi Taq polymerase (Promega)
Reaction volume: 20µl
For CAPS analysis an additional 5µl of a mix consisting of 2.5ml of 10x NEB buffer 4 and 0.5µl of BspHI (10U/µl; NEB) was added once the PCR was completed and the tubes incubated at 37° for 1 hour.
Expected products and validation
The CAPS product was separated on a 2.5% (w/v) agarose gel (Fig.1). csSr2 was tested on 122 diverse wheat lines from around the world including 78 Australian past and present cultivars (Table 1). The marker is associated with the presence or absence of the gene in 95% of the lines tested. Its accuracy increased to 100% in lines that were considered to lack Sr2. Potential discrepancies were observed in a few lines that were predicted to carry Sr2 but failed to show the CAPS marker. It is possible that some lines presumed to carry Sr2, were wrongly classified and lack the gene and that these lines were correctly identified as such by the marker. It is also not uncommon that wheat varieties released by breeders consist of mixtures and are heterogenous for important traits such as disease resistance or seed storage proteins. Alternatively, the marker associated with resistance may have mutated or recombined with the gene and is therefore not diagnostic for Sr2 in these lines. Future genetic studies of populations derived from these lines could clarify if the marker lacked accuracy or if wheat lines were wrongly classified.
Allele survey of diverse wheat germplasm with known (or presumed) Sr2 status using SSR marker gwm533 and CAPS marker csSr2 are available here: Sr2_gwm533.xlsx.
KASP markers for Sr2
Information kindly provided by Gina Brown Guedira.KASP
- SNP ID: wMAS000005
- Gene: Sr2
- Evidence: linked
- Primer Allele FAM: GTGCGAGACATCCAACACTCAC
- Primer Allele VIC: GTGCGAGACATCCAACACTCAT
- Primer Common: CTCAAATGGTCGAGCACAAGCTCTA
- FAM allele: G
- VIC allele: A
- FAM phenotype: susceptible
- VIC phenotype: resistant
This markers were developed based on the work of Mago et al. (4). For more information on KASP protocols, please check visit this link.
References
1. Identification and validation of markers linked to broad-spectrum stem rust resistance gene Sr2 in wheat (Triticum aestivum L.). Spielmeyer W, Sharp PJ, Lagudah ES. In: Crop Science, 2003, 43:333–336. DOI:10.2135/cropsci2003.0333.
2. Sequence tagged microsatellites for the Xgwm533 locus provide new diagnostic markers to select for the presence of stem rust resistance gene Sr2 in bread wheat (Triticum aestivum L.). Hayden MJ, Kuchel H, Chalmers KJ. In:Theoretical and Applied Genetics, 2004, 109:1641–1647. DOI:10.1007/s00122-004-1787-5.
3. BAC-derived markers for assaying the stem rust resistance gene, Sr2, in wheat breeding programs. McNeil MD, Kota R, Paux E, Dunn D, McLean R, Feuillet C, Li D, Kong X, Lagudah E, Zhang JC, Jia JZ, Spielmeyer W, Bellgard M, Appels R. In:Molecular Breeding, 2008, 22:15–24. DOI:10.1007/s11032-007-9152-4.
4. An accurate DNA marker assay for stem rust resistance gene Sr2 in wheat. Mago R, Brown-Guedira G, Dreisigacker S, Breen J, Jin Y, Singh R, Appels R, S. Lagudah ES, Ellis J, Spielmeyer W. In: Theoretical and Applied Genetics, 2011, 122:735-744. DOI:0.1007/s00122-010-1482-7.
