The concentration of yellow carotenoid pigments in durum wheat grain is an important quality criterion and is determined both by their accumulation during grain development and by their degradation by lipoxygenase enzymes (Lpx loci) during pasta processing (1). A deletion of one of the duplicated copies at the Lpx-B1 locus (Lpx1.1) was associated with a 4.5-fold reduction in lipoxygenase activity and improved pasta color (P<0.0001) but not semolina color, suggesting reduced pigment degradation during pasta processing (2). A molecular marker for the deletion was mapped on chromosome 4B in a population obtained from the cross between durum line UC1113 and variety Kofa. The variety Kofa carries the deleted Lpx1.1 allele contributing to its excellent pasta color. The markers presented here are useful to select for this deletion in durum wheat breeding efforts aimed to improve pasta color.
PCR-based marker for Lpx1.1 deletion in pasta wheat
Primer sequences
LOXA-L1 5’- CTGATCGACGTCAACAAC - 3’
LOXA-R1 5’- CAGGTACTCGCTCACGTA - 3’
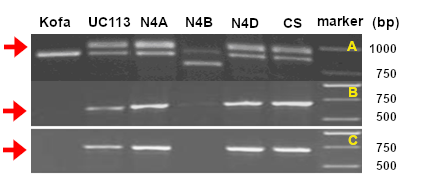
These primers amplify two bands in wild type durum, but the top band (Lpx1.1) is missing in Kofa (2). Three additional primers can be used in different combinations to confirm the Lpx-1.1 deletion (1):
LOXA-4BSL 5’- CTTACCTAGTACAATGTACTCTCTCT -3’
LOXA-4BSR 5’- ATGATGGTCTGGATCTGG -3’
LOXA-4BSL2 5’- TCTCATCAAGAGGTACCTACTTA -3’
Final concentrations of the reagents used in the PCR amplification
- 75-150 ng template DNA
- 100 nM of each primer
- 1 unit of Taq DNA polymerase
- 1.5 mM MgCl2
- 2.5 µl 10X buffer
- 200 mM dNTPs
Total volume: 25 µl reaction
PCR conditions (times for each step might vary according to the thermocycler model):
- Denaturing step: 3 minutes at 94°C.
- 5 touchdown cycles (-1°C each) [30 sec at 94 °C, 30 sec at 60–55 °C and 45 sec at 72 °C ].
- Amplification step (35 cycles)
- 94°C, 45 sec
- 55°C, 45 sec
- 72°C, 45 sec
- Extension step: 10 minutes at 72°C.
Expected products
Following amplification, the products are separated in 1% agarose gels at 65 V constant voltage.
References
1. Association of a lipoxygenase locus, Lpx-B1, with variation in lipoxygenase activity in durum wheat seeds. Hessler TG, Thomson MJ, Benscher D, Nachit MM, Sorrells ME. In: Crop Science, 2002, 42:1695-1700. DOI:10.2135/cropsci2002.1695
2. A deletion at the Lpx-B1 locus is associated with low lipoxygenase activity and improved pasta color in durum wheat (Triticum turgidum ssp. durum). Carrera A, Echenique V, Zhang W, Helguera M, Manthey F, Schrager A, Picca A, Cervigni G, Dubcovsky J. In: Journal of Cereal Science, 2007, 45:67-77. DOI:10.1016/j.jcs.2006.07.001
